Researchers from the University of Michigan (U-M) Medical School have found culture-independent evidence that the lung microbiome is enriched with gut bacteria, both in a murine model of sepsis and in patients with ARDS. In more severely critically ill patients, lung bacteria were more outnumbered by the misplaced gut bacteria. The researchers conclude that critical illness that involves the lungs has more to do with disruptions to the body’s natural population of microbes, or microbiome than previously thought. Their findings, just published in Nature Microbiology suggest that there may be a shared mechanism of pathogenesis in sepsis and ARDS.
The researchers performed animal and human studies. They saw the effect in mice with sepsis, and in 68 human patients with Acute Respiratory Distress Syndrome (ARDS). They studied the lung microbiome of patients with ARDS using special genetic tools and bacterial culture techniques, and compared them to samples from healthy volunteers.
Lead author Robert P. Dickson, MD, Assistant Professor in Internal Medicine, Division of Pulmonary & Critical Care Medicine and associate director of U-M's Center for Integrative Research in Critical Care, commented in a statement that previously most attempts to treat these critical illnesses have been aimed at fixing the disordered inflammation and tissue injury. This study raises the possibility that this inflammation and injury may actually be downstream consequences of an upstream source: disordered bacterial communities in the gut and lung, he added.
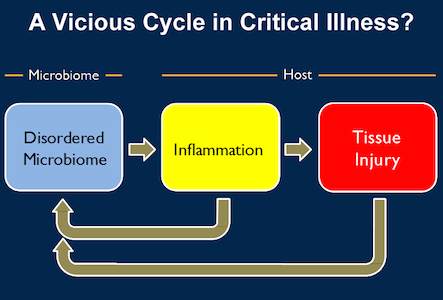
Dickson and his co-researchers believe that patients with these conditions may be stuck in a vicious cycle caused by dysbiosis or microbial imbalance. They suggest that the insult that caused the patient's or animal's illness in the first place triggers a "chicken and the egg" feedback loop.
Changes in the microbiome lead to inflammation, as the body's immune system tries to fend off what it sees as invaders. The inflammation in turn injures the lung tissue, and the injury and inflammation change the environment within the lung, allowing microbes that don't normally grow there to invade or to "bloom" if they were already present in low levels.
 To break the cycle and improve survival of critically ill patients research needs to focus on how to keep the microbiome relatively normal, starting with finding out where the gut bacteria in the lung came from originally. The researchers discovered that the upper respiratory tract was not the route in their animal study.
To break the cycle and improve survival of critically ill patients research needs to focus on how to keep the microbiome relatively normal, starting with finding out where the gut bacteria in the lung came from originally. The researchers discovered that the upper respiratory tract was not the route in their animal study.Possible explanations include “leaky” intestinal walls or gut bacteria being present in the lungs all along. Dickson noted that researchers have only recently started to think of the lungs as an ecosystem.
“We're just now sorting out the rules for how these bacterial communities get established, both in health and in critical illness”, said Dickson.
Next Steps
"In the long run, we need to start thinking of the microbiome as an organ that can fail in critically ill patients," commented Dickson. "We're studying how it gets disordered, how it impacts other organs, and how we can fix it. The importance of the microbiome in the ICU has been clear for decades, but with these new tools we're finally able to ask and answer the right questions. It's a really exciting time."
Dickson explained in an email to ICU Management & Practice that the team is currently studying the lung microbiome in sepsis and ARDS both at the lab bench and in the intensive care unit (ICU). In animal models of critical illness they are working to determine how these gut bacteria overtake the lung microbiome and trying to determine to what degree this change in the lung microbiome is a cause of inflammation and lung injury, or just a secondary effect of critical illness and its treatment.
In the ICU, the team is studying how common and clinically significant this alteration in the lung microbiome is.
"We want to know how commonly it occurs, what clinical exposure predict it, and whether it predicts the development of ARDS. Ultimately, we'd love to know if it's a feature of disease pathogenesis that can be targeted therapeutically,” said Dickson.
"To determine if a specific pathophysiological process is occurring, like translocation of gut bacteria to the lung, we should be able to learn something from a few dozen patients, so long as both their disease and their microbiota are well characterised. To do this right, we will need to sample multiple bacterial communities from each patient (gut, lung, pharynx, endotracheal tube) and collect longitudinal specimens."
"To understand specific interactions between lung bacteria and the host, our studies will need to be much larger. The microbiome is every bit as complex as the host's genome, and is more dynamic and variable between individuals. So I suspect we will need to be thinking like our colleagues in Genetics when it is time to power those studies."
Asked about potential treatments, Dickson noted that there have been several promising but modest studies of probiotics in the prevention of ventilator-associated pneumonia, adding: "I think we are really in the dark ages regarding if and how they work, or if they are relevant to ARDS."
"We don't know - in humans or even in animal models - whether giving probiotics via the mouth or a feeding tube alters lung microbiota. We don't know how the lung microbiome is altered by current oral hygiene practices (like chlorhexadine). It is conceivable that the lung and gut microbiomes will one day be therapeutically manipulated in the ICU (as with faecal microbiota transplants for recurrent C. difficile infections), but there is a lot we need to learn between now and then."
Sources: University of Michigan Health System; interview
Image credit: Top image, Flickr, CC by 2.0; middle image, credit University of Michigan
References:
Dickson RP, Singer BH, Newstead MW, Falkowski NR, Erb-Downward JR, Standiford TJ, Huffnagle GB (2016) Enrichment of the lung microbiome with gut bacteria in sepsis and the acute respiratory distress syndrome. Nature Microbiology, doi: 10.1038/nmicrobiol.2016.113
Latest Articles
ARDS, Sepsis, microbiome, how do gut bacteria get into lungs?
Researchers from the University of Michigan Medical School have found culture-independent evidence that the lung microbiome is enriched with gut bacteria both in a murine model of sepsis and in patients with ARDS. In more severely critically ill patients,