ICU Management & Practice, Volume 23 - Issue 1, 2023
Abstract
Sepsis and antimicrobial resistance are two major public health problems. Sepsis is defined as a dysregulated response of the body to a bacterial infection with organ dysfunction. The 2021 Surviving Sepsis Campaign SSC guidelines proposed characterising the infection based on clinical assessment and the dysregulated host response using clinical scores. In children, the sepsis definition is based on the Sepsis-2 definition for adults using SIRS criteria and the Pediatric Logistic Organ Dysfunction (PELOD) score. Based on clinical signs (fever, blood pressure, respiratory rate, mental status…) and a dysregulated host response, this approach has a high sensitivity to detect sepsis and helps initiate antibiotic therapy rapidly. This approach, however, lacks specificity, resulting in numerous unnecessary treatments with broad-spectrum antibiotics, contributing to antimicrobial resistance. Biomarkers of sepsis such as circulating blood leukocytes, C-reactive protein (CRP), procalcitonin (PCT), lactate, IL-6, presepsin, Pancreatic Stone Protein (PSP), transcriptomic profiling, etc., did not improve the performance of the clinical approach and are usually not recommended in guidelines.
An ideal biomarker to detect bacterial infection and sepsis should therefore have a sensitivity as high as that of the clinical approach but with better specificity and a sufficiently high negative predictive value to rule out infection and sepsis. It should improve diagnostic accuracy when combined with the clinical approach. Ideally, a useful biomarker should detect sepsis even before clinical manifestations and could be used to monitor hospitalised patients at high risk of nosocomial sepsis. Such ability for a pre-symptomatic diagnosis of nosocomial sepsis may help in early sepsis identification, control of the source of sepsis and rapid initiation of antibiotics. It should also follow the course of sepsis resolution and help antibiotic de-escalation, as proposed with PCT. Finally, this marker should ideally be easily available in all hospitals, rapid and simple to measure, robust and affordable.
We propose a checklist of criteria to assess the usefulness of a biomarker to be integrated into sepsis guidelines.
Introduction
Infectious diseases are a major public health problem throughout the world. Bacterial infections can be complicated by sepsis, a dysregulated response of the host to an infection associated with organ dysfunction (Singer et al. 2016). Septic shock is a subset of sepsis, with circulatory collapse and metabolic dysfunction associated with high mortality. Worldwide, 48.9 million people develop sepsis each year, and 11 million die of septic shock (Rudd et al. 2020). Sepsis and septic shock represent the second cause of mortality worldwide, behind cardiovascular diseases, and the leading cause of death in children (WHO 2018). The diagnosis and management of sepsis are based on criteria and algorithms published in international guidelines (Evans et al. 2021).
The definition of sepsis has evolved over time. In 1992, in the first set of definitions, sepsis was defined as a suspected bacterial infection associated with a systemic inflammatory response syndrome (SIRS) (Bone et al. 1992). In 2003, sepsis was defined as a suspected bacterial definition associated with a more complete SIRS definition (Levy et al. 2016). In 2016, a third set of definitions of sepsis was proposed: sepsis was defined as a bacterial infection causing organ dysfunction (Singer et al. 2016). The concept of SIRS was abandoned. Since October 2021, the latest set of sepsis definitions was proposed by the Surviving Sepsis Campaign (SSC) (Evans et al. 2021). Sepsis was defined as a bacterial infection associated with a dysregulated response of the body with SIRS and organ dysfunction. Guidelines for the management of sepsis were first proposed by the SSC in 2004 (Dellinger et al. 2004); they have since been revised regularly, with the latest update published in October 2021 (Evans et al. 2021). The key objectives of these SSC guidelines are a rapid sepsis diagnosis to initiate appropriate treatment within 45 to 60 minutes, in particular antibiotics and source control, to avoid progression to septic shock, and to reduce sepsis-associated morbidity and mortality (Evans et al. 2021). In children, the sepsis definitions currently used were proposed in 2005 and are similar to those proposed for adults (Sepsis-2), however, including the 1992 SIRS criteria (Goldstein et al. 2005). The SSC management guidelines for sepsis in children were published in 2020 (Weiss et al. 2020).
The Diagnosis of Sepsis
In adults and in children, the diagnosis of sepsis is based on an early diagnosis of a bacterial infection and the identification of a dysregulated response of the body with organ dysfunction (Evans et al. 2021; Weiss et al. 2020). There are, therefore, two different diagnostic approaches to be made in parallel. There are also two different therapeutic approaches to be taken in parallel: the control of infection with antimicrobials and the source of infection, and the treatment of the dysregulated response of the body and organ dysfunction, with volume resuscitation, organ support, and adjuvant therapies.
Diagnosis of (suspected) infection
In the case of (suspected) sepsis, it is recommended to confirm or rule out bacterial infection and to continuously reassess the infectious diagnosis, to initiate, modify or stop antibiotic therapy (Evans et al. 2021). Unfortunately, bacterial cultures, the gold standard to diagnose bacterial infections, take between 24 and 48 hours to give meaningful results to the clinician and do not help to make decisions as to start antibiotics or not. They may also be false negatives due to prior antibiotic therapy or inadequate sampling. It is, however, a strong recommendation in the sepsis management guidelines that antibiotic therapy should be initiated as soon as possible, at least less than 45 to 60 minutes after suspicion of sepsis (Evans et al. 2021). Initiating broad-spectrum antibiotics in all patients with suspected sepsis will result in unnecessary treatments in 60 to 70% of patients who end up not having sepsis, in part because signs are similar between severe viral and bacterial infections and also with severe inflammatory processes (Klein Klouwenberg 2015). This is problematic since antibiotics are associated with secondary toxic effects and will contribute to the development of antibacterial resistance both in the patient and in the ICU. Overtreatment with antibiotics has been particularly problematic during the COVID-19 pandemic (Lai et al. 2021; Ghosh et al. 2021). It has recently been estimated that 4.95 million (3.62–6.57) deaths were yearly associated worldwide with antimicrobial resistance (Antimicrobial Resistance Collaborators 2022).
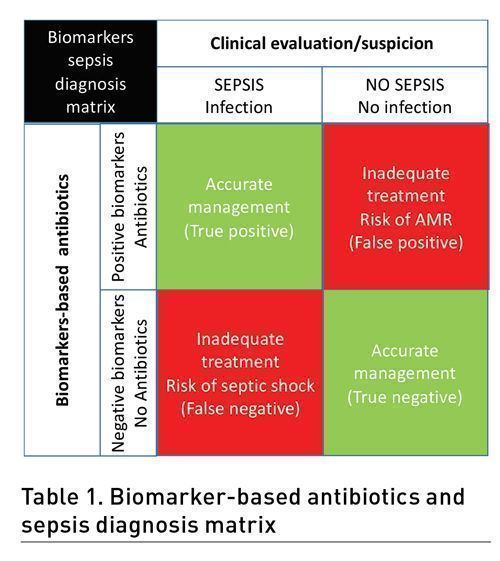
Biomarkers have been extensively studied to help diagnose rapid bacterial infection in patients with suspected sepsis. They usually do not help to increase sensitivity since sensitivity is already very high with clinical signs and scores. However, they may be used to increase specificity to restrict antibiotic treatment only to patients with a bacterial infection. In addition, they should have a good negative predictive value to rule out a bacterial infection in patients with suspected sepsis and prevent unnecessary antibiotic treatments. Therefore, we believe biomarkers may be a useful addition to the current diagnosis based on clinical signs and scores. This matrix presented in Table 1 describes four clinical conditions: adequately treated sepsis, inadequate sepsis treatment at risk for aggravation, unnecessary antibiotic therapy at risk of bacterial resistance, and adequate no (or suspended) antibiotic therapy.
Once antibiotic therapy has been initiated, it should be reassessed daily (pursue, modify or stop) based on the results of bacterial cultures and clinical evolution (Evans et al. 2021). The goal is to tailor antibiotic therapy to a given patient with a given infection. Such an approach is likely to shorten the duration of antibiotic treatment, but still frequently based on fixed duration. Biomarkers may help the clinician fulfil this goal and shorten the duration of antibiotic treatment. Daily measurement of PCT is used to guide the clinician to de-escalate and stop antibiotic therapy in adults and children. A similar approach using CRP has also been used but not recommended (Evans et al. 2021). However, the level of evidence remains low (Evans et al. 2021; Weiss et al. 2020).
Diagnosis of dysregulated response of the body
The diagnosis of dysregulated response of the body is based on SIRS criteria and/or organ dysfunction according to the SSC 2021 guidelines (Evans et al. 2021). SIRS criteria are those originally described in 1992 for adults and children (Bone et al. 1992). They evaluate three non-specific clinical parameters: temperature, heart rate, and respiratory rate, as well as two laboratory results: circulating blood leukocytes and PaCO2, which require an arterial puncture and access to a blood gas analyser. The dysregulated organ response is based on two clinical scores, the National Early Warning Score (NEWS) and the Modified Early Warning Score (MEWS) (strong recommendation, moderate quality evidence) (Evans et al. 2021). The NEWS evaluates eight non-specific clinical parameters and three organs (lung, circulation/heart, and brain), and the MEWS evaluates six non-specific clinical parameters and the same three organs. These two scores do not require blood sampling. NEWS and MEWS are not used in children. The gold standard for the dysregulated organ response is the Sequential Organ Failure Assessment (SOFA) score which assesses the function of six organs impacted by sepsis (lung, circulation/heart, brain, liver, kidney, coagulation), combining five clinical parameters and four laboratory results. This score is rather complex and requires venous and arterial blood tests and requires time to calculate - about two hours. The qSOFA (quick SOFA) is a simple and fast version of the SOFA which takes < 5 minutes. qSOFA was introduced in 2016, measuring only three non-specific clinical parameters. However, the 2021 SCC guideline suggested that the qSOFA should no longer be used (Evans et al. 2021). In paediatrics, the organ failure score is the Pediatric Logistic Organ Dysfunction (PELOD) score which assesses the function of six organs (Goldstein et al. 2005). The lactate assay is proposed in the 2016 Sepsis-3 definition. It is part of the first hour of management of sepsis (Singer et al. 2016), but it is not sensitive nor specific enough to make the early diagnosis of sepsis and dysregulated organ response. In 2021, lactate measurement was only weakly recommended with a low degree of evidence. Lactate is proposed to be used as an additional assay to guide volume resuscitation in septic shock with again a low recommendation and low level of evidence (Evans et al. 2021). The lactate assay is often used in paediatrics but is not recommended by the 2020 SSC guidelines (Weiss et al. 2020).
Clinical Signs, Scores, and Biomarker
The consequence of the four different definitions of sepsis since 1992, the absence of a gold standard specific test for early diagnosis of sepsis and no sepsis, and the change in clinical scores to be used have led to confusion among clinicians. More than half of the clinicians (53.8%) pragmatically use a mixture of all these scores (Ventura 2021); 89.7% measure circulating blood leucocytes, 92.3% CRP, 84.6% PCT, and 100% lactate in case of suspicion of sepsis. Only 35.9% use the Sepsis-3 definition alone, 34.2% still calculate the qSOFA, and 44.7% use the complete SOFA score. This indicates that in daily practice, clinicians use more biomarkers than scores. As previously mentioned, the use of biomarkers should primarily help to rule out the diagnosis of (suspected) infection and sepsis, while most biomarker studies seek to show their utility in diagnosing (suspected) infection and sepsis. The ideal biomarker of infection and sepsis should therefore have a sensitivity similar to the non-specific, highly sensitive clinical approach but should carry a better specificity, with a very good negative predictive value to rule out infection and sepsis. Ideally, biomarkers should be able to detect sepsis even before clinical suspicion, which is only possible by routine monitoring (daily measurements) in hospitalised patients at high risk for nosocomial sepsis. This is the concept of pre-symptomatic diagnosis of nosocomial sepsis. Finally, a biomarker should be able to help with the decision of antibiotic de-escalation, coupled with clinical evaluation.
In summary, one or more biomarkers should help in the diagnosis of sepsis and nosocomial sepsis in the three chronological phases: the pre-sepsis or pre-symptomatic phase (with daily monitoring of hospitalised patients at risk), the clinical onset of sepsis phase, and the post-sepsis phase (Table 2).
Discussion
Many biomarkers of sepsis have been studied. A recent review identified 258 sepsis biomarkers (Pierrakos et al. 2021). Already in 2009, the International Sepsis Forum Colloquium on Biomarkers of Sepsis proposed to develop a systematic framework for identifying and validating biomarkers of sepsis and to promote collaboration between investigators, the biomarkers industry, and regulatory agencies (Marshall and Reinhart 2009). Unfortunately, the numerous studies carried out since then have remained rather heterogeneous and have not made it possible to identify and compare accurately one or more sepsis biomarkers with the capacity to screen, diagnose, stratify risk, monitor, and use them as a surrogate endpoint. In 2020, Prof. John Marshall and Prof. Jean-Louis Vincent concluded: “Continuing to produce reports of novel biomarkers without developing a more rigorous framework to evaluate them and establishing a recognised purpose is futile: it is time for a reappraisal of the possible roles of biomarkers in sepsis” (Pierrakos et al. 2020).
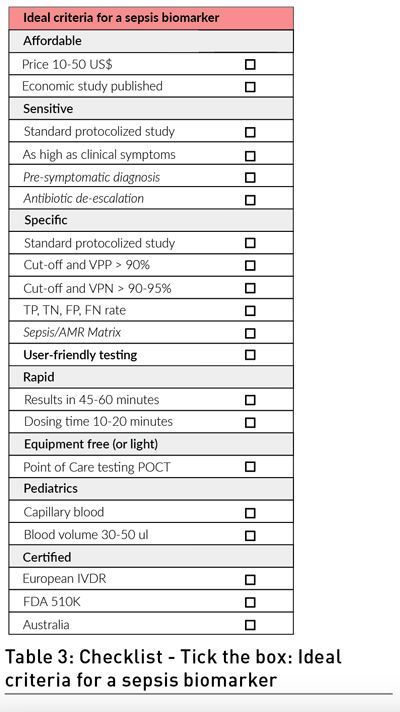
It is fundamental from now on to carry out future clinical studies on sepsis biomarkers analysing their specificity (PPV and NPV with cut-offs), their capacity to make a pre-symptomatic diagnosis, and their potential in the de-escalation of antibiotics. We, therefore, propose to establish a checklist (Table 3) with the basic requirements that biomarkers of sepsis should meet and a standard protocol for biomarker sepsis studies so that future studies can be comparable and can answer the urgent questions raised by the major public health problems of sepsis and antimicrobial resistance. This standard protocol should include three specific protocols according to the three phases of sepsis (pre-sepsis, sepsis, post-sepsis) the studies would like to investigate.
The first and very important common points for all biomarkers standard protocols are the definition of infection and the definition of sepsis used. The diagnosis of infection can be confirmed by bacteriological culture results in only 40-50% of cases (Klein Klouwenberg et al. 2015). In this context, confirmation of the diagnosis of infection in a biomarker study must be reviewed by an adjudication committee, independentof the investigators, based on the analysis of several parameters and patient charts. The most detailed and recommended way is to use the Centers for Disease Control (CDC) guidelines of January 2022 [20]. A simpler way, and the recommended minimum, is the one proposed in a 2017 biomarker study (García de Guadiana-Romualdo et al. 2017). It is proposed to consider two ways to confirm infection (Singer et al. 2016). A clinically documented infection (CDI) occurs when there are clinical signs or symptoms and radiological or/and analytical findings of infection but without microbiological proof (Rudd et al. 2020). The microbiologically documented infection (MDI) includes bacteraemia (microbiologically documented infection with bacteraemia) and microbiologically documented local infection without positive blood cultures. In adults, it is, therefore, necessary to diagnose infection as described above, and the SIRS-1992, NEWS, MEWS, and SOFA scores (PELOD score for paediatric) must be calculated for each biomarker determination. The following classical biomarkers CRP, procalcitonin, circulating blood leukocytes, and serum lactate, must be measured for each new biomarker determination and the biomarkers used to calculate the SIRS-1992, the SOFA or PELOD scores.
From a statistical viewpoint, it is necessary to analyse the specificity, sensitivity, AUC, ROC curves, positive and negative predictive values, with cut-off values of classical and new biomarkers. The aim of such a standardised observational study is to answer whether the administration or not of antibiotics, the performance or not of complementary examinations, a hospitalisation or a return to home, the combination biomarkers, and eventually, a clinical score integrating a biomarker are possible. Ideally, an economic study should be carried out in parallel to determine the cost of the biomarker assay and the savings made (antibiotics, complementary examinations, length of stay, etc.). The objective of these observational protocolised and standardised studies must be able to carry out future interventional impact studies that could evaluate the real benefits of sepsis biomarkers in terms of incidence, prevalence, antibiotic use, morbidity, and mortality related to sepsis and antimicrobial resistance.
Biomarker studies in the pre-sepsis phase require more resources because all the common points must be analysed every day or even several times a day. Statistical analyses are more complicated, especially for analysing the kinetics with absolute and relative daily changes in biomarkers to predict infection and/or nosocomial sepsis.
Biomarkers studies in the sepsis phase require to determine all the common parameters at once as soon as sepsis is clinically suspected. Time parameters, such as the time between suspected sepsis, biomarker results and antibiotic administration, must be studied and considered.
A biomarkers study in the post-sepsis phase requires biomarkers daily dosing after the diagnosis of sepsis to determine the effectiveness of treatment, early detection of recurrence, and the duration of antibiotic therapy (de-escalation). This type of study also requires an adjudication committee to determine the adequacy of antibiotic treatment.
The study of combinations of different biomarkers could improve the sensitivity and specificity of a single biomarker. It might be interesting to develop a clinical pre-test probability score for each of these biomarkers to further improve sensitivity and specificity.
Conclusion
In the context of the global public health emergency to solve the problems of sepsis and antimicrobial resistance, we propose to standardise sepsis biomarker studies to determine which of the classic and new biomarkers might be the ideal biomarker(s) to confirm or exclude infection and sepsis, to pre-symptomatically detect nosocomial infection and sepsis, and to help guide antibiotic de-escalation.
Both classical sepsis and infection biomarkers (circulating blood leukocytes, CRP, PCT and lactate) and new biomarkers (e.g., interleukin IL-6, presepsin, PSP, transcriptomic profiling, etc.) should therefore be studied or re-studied using a standardised approach to determine which biomarker(s) answer(s) clinicians' questions. It will also be necessary that the technology for determining the ideal sepsis biomarker(s) can be easily transferred from research laboratories to clinical use and meet regulatory criteria such as FDA 510(k), the European In Vitro Diagnostic Regulation IVDR2022, etc.
In addition, and this is mandatory, the biomarker result should be obtainable within 45 to 60 minutes to initiate (or not) antibiotic therapy quickly, as required by the SSC 2021 guidelines (Evans et al. 2021) for adults and the SSC 2020 for children (Weiss et al. 2020). Probably only a point-of-care testing POCT, with a 10-20 minute dosing time, can allow such a rapid result, and it must be able to fulfil to a large extent the ASSURE criteria (Affordable, Sensitive, Specific, User-friendly, Rapid, Equipment free) for a sepsis diagnosis test (Bissonnette and Bergeron 2010). In children, especially neonates, a capillary sample and less than 30-50 µl of blood volume would be ideal. Economic studies should also be able to determine the financial consequences of sepsis biomarker testing on public health costs, and the price of the biomarker assay is not expected to exceed USD 10 to 50. The objective of all these studies could allow one or several infection and sepsis biomarkers that tick all the boxes on the checklist (Table 3) to be included in the next adults and paediatrics international Surviving Sepsis Campaign guidelines to help solve the major problems of sepsis and antimicrobial resistance.
Disclaimer
Point-of-View articles are the sole opinion of the author(s) and they are part of the ICU Management & Practice Corporate Engagement or Educational Community Programme.
References:
Antimicrobial Resistance Collaborators (2022) Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. Lancet. 399(10325):629-655.
Bissonnette L, Bergeron MG (2010) Diagnosing infections--current and anticipated technologies for point-of-care diagnostics and home-based testing. Clin Microbiol Infect. 16(8):1044-53.
Bone RC, Balk RA, Cerra FB et al. (1992) Definitions for sepsis and organ failure and guidelines for the use of innovative therapies in sepsis. The ACCP/SCCM Consensus Conference Committee. American College of Chest Physicians/Society of Critical Care Medicine. Chest. 101(6):1644-55.
CDC/NHSN Surveillance Definitions for Specific Types of Infections. Available at
https://www.cdc.gov/nhsn/pdfs/pscmanual/17pscnosinfdef_current.pdf
Dellinger RP, Carlet JM, Masur H et al. (2004) Surviving Sepsis Campaign guidelines for management of severe sepsis and septic shock. Crit Care Med. 32(3):858-73.
Evans L, Rhodes A, Alhazzani W et al. (2021) Surviving Sepsis Campaign: International Guidelines for Management of Sepsis and Septic Shock 2021. Critical Care Medicine. 49(11):e1063-e1143.
García de Guadiana-Romualdo L, Berger M, Jiménez-Santos E et al. (2017) Pancreatic stone protein and soluble CD25 for infection and sepsis in an emergency department. Eur J Clin Invest. 47(4):297-304.
Ghosh S, Bornman C, Zafer MM (2021) Antimicrobial Resistance Threats in the emerging COVID-19 pandemic: Where do we stand? J Infect Public Health. 14(5):555-560.
Goldstein B, Giroir B, Randolph A (2005) International Consensus Conference on Pediatric Sepsis. International pediatric sepsis consensus conference: definitions for sepsis and organ dysfunction in pediatrics. Pediatr Crit Care Med. 6(1):2-8.
Klein Klouwenberg PM, Cremer OL, van Vught LA et al. (2015) Likelihood of infection in patients with presumed sepsis at the time of intensive care unit admission: a cohort study. Crit Care. 19(1):319.
Lai CC, Chen SY, Ko WC, Hsueh PR (2021) Increased antimicrobial resistance during the COVID-19 pandemic. Int J Antimicrob Agents. 57(4):106324.
Levy MM, Fink MP, Marshall JC et al. (2003) SCCM/ESICM/ACCP/ATS/SIS. 2001 SCCM/ESICM/ACCP/ATS/SIS International Sepsis Definitions Conference. Crit Care Med. 31(4):1250-6.
Marshall JC, Reinhart K; International Sepsis Forum (2009) Biomarkers of sepsis. Crit Care Med. 37(7):2290-8.
Pierrakos C, Velissaris D, Bisdorff M et al. (2020) Biomarkers of sepsis: time for a reappraisal. Crit Care. 24(1):287.
Rudd KE, Johnson SC, Agesa KM et al. (2020) Global, regional, and national sepsis incidence and mortality, 1990-2017: analysis for the Global Burden of Disease Study. Lancet. 395(10219):200-211.
Singer M, Deutschman CS, Seymour CW et al. (2016) The Third International Consensus Definitions for Sepsis and Septic Shock (Sepsis-3). JAMA. 315(8):801-10.
Ventura F (2021) Diagnosis of Sepsis. chez Area Critica, Bologna, 2021.
Weiss SL, Peters MJ, Alhazzani W et al. (2020) Surviving Sepsis Campaign International Guidelines for the Management of Septic Shock and Sepsis-Associated Organ Dysfunction in Children. Pediatr Crit Care Med. 21(2):e52-e106.
WHO (2020) The top 10 causes of death. Available at https://www.who.int/news-room/fact-sheets/detail/the-top-10-causes-of-death