ICU Management & Practice, Volume 21 - Issue 5, 2021
A significant research gap exists in the field of the lung microbiome and pneumonia development in paediatric population. Its study may improve nosocomial pneumonia prevention and help to achieve zero pneumonia rates.
Most frequent hospital acquired infections (HAI) in Paediatric Intensive Care Units (PICU), are ventilator associated pneumonia (VAP), catheter-related urinary tract infection and catheter-related infection. All can be the onset of a sepsis or septic shock (Esteban et al. 2013; Goldstein et al. 2005).
What is the VAP Impact in Paediatric ICU?
The incidence of VAP is around 8-9% of ventilated adults and 14% of ventilated paediatric patients (Elward et al. 2002). Two universal criteria required for VAP diagnosis are PICU admission longer than 72 hours and intubation for more than 48 hours. In paediatrics, the criteria of the Centre for Disease Control are the most widely used, and include clinical-analytical, radiological, and microbiological criteria (Horan et al. 2008).
VAP has high morbidity and mortality in PICU, despite the pneumonia zero programmes. Risk factors for developing VAP depend on the patient and prevention measures conducted. The mechanism for VAP apparition are described in three pathophysiological ways: oropharyngeal or stomach secretions aspiration; direct inoculation through the endotracheal tube (ETT); and less frequently through haematogenous dissemination. Preventive measures are targeted against these principles but it’s difficult to achieve zero pneumonia rates, especially in paediatric settings (Branch-Elliman et al. 2015; Nair et al. 2015).
Patients with VAP are especially sensible for infections caused by antimicrobial resistance microorganism (AMR), due to patient factors (comorbidities, broad-spectrum antibiotics) as well as external factors (invasive devices, long-term hospitalisation). AMR, and especially multi drug resistance (MDR), are an important public health issue because of the severity of the infections they may cause, the difficulty to establish a correct empiric treatment, the ability for spreading MDR and the absence of new antibiotics against those pathogens (Boucher et al. 2009; Marston et al. 2016; Nathan et al. 2014).
National actions have proposed strategies to prevent VAP and MDR, as the ENVIN_HELICS register or “Pneumonia zero” programme. National PICUs have collaborated with these projects in order to promote and reinforce the safety culture in the PICU of the National Health System (Grau et al. 2013; Martínez-Martínez et al. 2010; Jordan et al. 2016). Results of these programmes have detected relatively high rates of VAP in paediatrics (5-7 VAP/1000 days VM) (Jordan et al. 2014).
Are There Any Preventive Risk Factors?
Lowest age, previous comorbidities, patient severity, and ETT length are the main risk factors, but most of these are not susceptible to be changed. Strategies to prevent rates follow international bundles, like general strategies (airway manipulation, hand cleansing, use of cuffed tubes -pressure around 20 cmH2O-, head of the bed at 30-45º) similar than in adults. However, other recommended measures cannot be followed in children, such as the subglottic secretions aspiration (there is no dispositive for paediatrics), or digestive tube selective decontamination (controversial in children).
Due to the high rates of VAP in paediatric patients and the difficulties in the capacity to modify intrinsic characteristic of the patients, new approaches to this nosocomial infection are required.
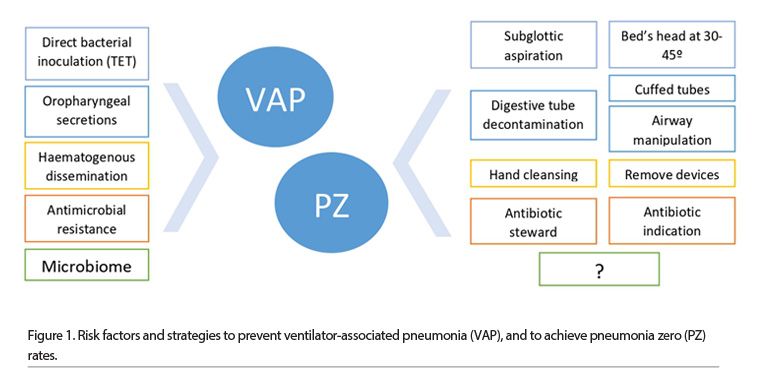
Attending to the principal pathophysiological ways of VAP development, oropharyngeal colonisation may be one of the investigation attention points. Either if the respiratory infection is secondary to the oropharyngeal aspirations or by direct inoculation, the microorganism responsible for VAP is commonly one of those colonising the airway. It has been defined that colonisation of the upper airway may be the beginning of the process that can conduce to a tracheobronchitis to even to a VAP. But better understanding might explain why patients develop one of these infections. Bacterial Microbiome (BM) could have an explanation for some of this concern in VAP (Pouline et al. 2017). Figure 1 shows risk factors and strategies to prevent VAP.
What is Human Microbiome?
The human microbiome is defined as the ecological community of commensal, symbiotic and pathogenic microorganisms found in humans. Microbiota has been found to be crucial for immunologic, hormonal and metabolic homeostasis of their host (The Human Microbiome Project Consortium 2012). It has been defined as a new factor to develop severe community infections and VAP in paediatrics. Not only are the main bacterial microorganism detected on traditional cultures, but also other bacterial commensals (Mourani et al. 2021). A great advantage of Next Generation Sequencing (NGS) analysis for BM detection is its capability to identify hard-to-grow or even uncultivable bacteria, so it would be useful to find another microorganism implicated in VAP development. Even BM could be a risk factor for bacterial resistance.
How Can We Analyse the Microbiome?
The use of next-generation sequencing and multiomic analysis reveals new insights on the identity of microbes in the lower airways blurring the lines between commensals and pathogens. Microbes are not found in isolation; rather they form complex meta-communities where microbe-host and microbe-microbe interactions play important roles on the host susceptibility to pathogens. In addition, the lower airway microbiota exerts significant effects on host immune tone (Wu et al. 2018).
In order to analyse it, there are two types of procedures: metagenomics and analysis of BM. The first relies on the sequencing of all nucleic acid sequences obtained in a sample. It allows to detect a new species in a sample. BM analysis is based on sequencing of specific regions of the DNA genes encoding the 16S ribosomal RNA (rRNA) subunit (Clarridge et al. 2004). This is possible by the presence of regions that are conserved throughout the bacterial kingdom within these genes, which allows PCR amplification using universal primers and permits the determination of bacterial composition in samples.
What is the Role of Microbiome in VAP Development in Children?
Data about mouth/nasopharyngeal (NF)/tracheal microbiome in paediatrics is scarce, but promising. Respiratory mucosa is immediately colonised after birth, by mother’s bacteria. The colonisation type may be different depending on genetic and environmental factors and changes during the first years of life. An equilibrium seems to exist between the invasive and commensal bacteria when the patient is healthy, with significantly higher BM diversity. In relation with caries, higher proportion of mouth Lactobacillus and anaerobic agents increase its infection, due to higher acids production (Tanner et al. 2018). In other paediatric pathology as bronchiolitis, four clusters of airway microbiota have been identified. Proportion of bronchiolitis was lowest in infants with Moraxella-dominant profile (14%) and highest in those with Staphylococcus-dominant profile (57%). By contrast, Corynebacterium/Dolosigranulum-dominant profile had low proportion of infants with bronchiolitis (17%) (Hasegawa et al. 2017). Other authors note that some viruses’ infections, such as influenza, may disrupt interactions between host microbial communities and host defence, thereby contributing to the pathogenesis of secondary bacterial infections (Hanada et al. 2018). In medium otitis the increasing trend in colonisation of otopathogen genera has also been correlated positively with frequencies of upper respiratory tract infection (Chonmaitree et al. 2017).
Two studies showed a decrease in bacterial diversity in the pulmonary microbiome with prolonged MV, with an increase in Pseudomonas and Acinetobacter species, even if another pathogen was causing the pneumonia (Mourani et al. 2021). Therefore, development of pneumonia could be possible due to selective pressure on the existing microbiome towards the selection of single bacteria species. One explanation is that commensal bacteria can release lipopolysaccharides and peptidoglycans which go through the oropharyngeal and respiratory epithelium. This capability is increased in inflammation situations, as commonly happens in critical patients. As an example, in children Respiratory Syncytial Virus has been determined to cause inflammation which conduces to Haemophilus clusters and responsible for a major severity of the viral infection (Rosas-Salazar et al. 2016; Zakharkina et al. 2017).
Longitudinal studies with daily cultures of endotracheal aspirates suggest that in the majority of the patients, the trachea is colonised with causative pathogen at least 1–2 days before pneumonia develops (Miyaki et al. 2005). However, colonisation does not frequently progress to pneumonia and traditional cultures need around three days to be positive. BM surveillance could also help in earlier treatment, which might allow for a shorter course of systemic antibiotics or even the use of inhaled antibiotics alone (Dickson et al. 2014; Kelly et al. 2016; Wexell et el. 2016).
If it was determined that children who develop VAP have different BM than children who do not, especially during the first admission days, some prevention actions could be implemented. It would be local or systemic measures, different than chlorhexidine or floral decontamination now recommended, as lactic inhibitors or anti-anaerobic products. Other decisions may be precociously done, such as antibiotic treatment in individualised cases.
Is There Other Microbiome of Non-Human Origin in Children?
Another focus related to HAI is surface contamination. There are contaminated surfaces in a hospital depending on the setting. In PICU most high-touch points are bed rails, supply cart/bedside table, computer mouse and intravenous pumps (Menis et al. 2011).
There are pathogens associated with each mode of transmission and environmental reservoir, especially multi-resistant microorganisms related with HAI. Environmental contamination may contribute to the transmission of healthcare pathogens when healthcare workers contaminate their hands or gloves by touching contaminated surfaces, or when patients come into direct contact with contaminated surfaces.
There are different methods for evaluating the cleaning of surfaces: the visual inspection (the information is not reliable and subjective); fluorescent marking (not specific, sensitive and the low cost); adenosine triphosphate control system (highly sensitive, not specific and expensive) (Nante et al. 2017); and microbiological analysis and colony counts (sensitive, specific, rapid and accurate but complicated) (White et al. 2007).
Since there are no scientific standards to measure the effect of an individual cleaner, or assess environmental cleanliness, finding the evidence to benefit the control of infection is further hampered. Analysis of microbiota samples of surfaces and medical devices allow the identification of pathogens considered possible reservoir in hospital areas and adoption of new cleaning methods and strategies for prevention and control infections.
What Do We Know From Our Paediatric Population?
Our group conducted a prospective preliminary study about microbiome and infectious disease in paediatric patients. Healthy subjects, cases with invasive pneumococcal disease and children with viral infection were analysed. The sample processing was the same as described in the methodology section, and microbiota was analysed by NGS. Results showed three different nasopharynx microbiota patterns. It was detected that Dolosigranulum genera seemed to have a protective profile and was significantly associated to healthy individuals. Second risk pattern was represented by Streptococcus, when it was detected together with a high diversity of anaerobic genera; it was associated with pneumococcal invasive disease. Third pattern was rich in Moraxella and Haemophilus and showed a trend to be related to viral respiratory infection.
What Are We Expecting in the Near Future?
1. A significant research gap exists in the study of the lung microbiome and pneumonia. General interest about human microbiome has been increasing during the last ten years. Scientific platforms have references of microbiota in humans since the 70’s but what has changed is the way that this microbiota is analysed, thanks to the NGS method. As previously reported, there have not been new explanations about why a patient develops VAP or not, even when patients have similar risk factors, pathologies or colonisation.
2. Complex microbial communities exist in the upper and lower airway. Microbe-host interactions blur the line between pathogen and commensal. If new sequencing methods were useful to diagnostic nasopharyngeal BM (NBM) pattern for VAP developing, a specific NBM panel would be defined and developed. This research may also allow translating the methodology to other causes of nosocomial infection, such as central line catheter associated infections or urinary tract infections associated with urinary catheter.
3. Regarding surface bacterial contamination, there is not much information, nor much about the method to be used for the diagnosis. To define how a PICU is colonised may be an opportunity to analyse if bacterial differences exist along time and if these colonising bacteria are the same that cause VAP. If surface microbiome results are useful, new panels for diagnosis should be developed depending on the bacterial species detected.
Conclusion
The microbial community of the lung may play an important role in pneumonia impacting susceptibility and the natural history of disease. Research interest, focused on determining the influence of NBM in the development of VAP in paediatric critically ill patients, and on analysing PICU surfaces BM colonisation, may improve VAP prevention. Added to previous VAP preventive measures, it may allow to reach zero pneumonia rates.
Conflict of Interest
None.
References:
Branch-Elliman W et al. (2015) Determining the ideal strategy for ventilator-associated pneumonia prevention: Cost-benefit analysis. Am J Respir Crit Care Med, 192:57-63.
Boucher HW et al. (2009) Bad bugs, no drugs: no ESKAPE! An update from the Infectious Diseases Society of America. Clin Infect Dis, 48(1):1-12.
Chonmaitree T et al. (2017) Nasopharyngeal microbiota in infants and changes during viral upper respiratory tract infection and AOM. PLoS ONE, 12(7):e0180630.
Clarridge JE (2014) Impact of 16S rRNA gene sequence analysis for identification of bacteria on clinical microbiology and infectious diseases. Clin Microbiol Rev, 17:840-62.
Dickson RP et al. (2014) Towards ecology of the lung: new conceptual models of pulmonary microbiology and pneumonia pathogenesis. Lancet Respir Med, 2:238-46.
Elward AM et al. (2002) Ventilator-associated pneumonia in pediatric intensive care unit patients: risk factors and outcomes. Pediatrics, 109(5):758-64.
Esteban E et al. (2013) The impact of a quality improvement intervention to reduce nosocomial infections in a PICU. Pediatr Crit Care Med,14(5):525-32.
Goldstein B et al. (2005) International paediatric sepsis consensus conference: definitions for sepsis and organ disfunction in paediatrics. Pediatr Crit Care Med, 6:2-8.
Grau S et al. (2013) How to measure and monitor antimicrobial consumption and resistance. Enferm Infecc Microbiol Clin, 31(Suppl 4):16-24.
Hanada S et al. (2018) Respiratory Viral Infection-Induced Microbiome Alterations and Secondary Bacterial Pneumonia. Front Immunol, 9:2640.
Hasegawa K et al. (2017) Nasal Airway Microbiota Profile and Severe Bronchiolitis in Infants. A Case-control Study. Pediatr Infect Dis J, 36:1044–1051.
Horan TC et al. (2008) CDC/NHSN surveillance definition of health care-associated infection and criteria for specific types of infections in the acute care setting. Am J Infect Control, 36(5):309-32.
Jordan I et al. (2014) A national multicentre study on nosocomial infections in PICU. An Pediatr (Barc), 80(1):28-33.
Jordan I et al. (2016) Trends in nosocomial infections and multidrug-resistant microorganisms in Spanish pediatric intensive care units. Enferm Infecc Microbiol Clin, 34(5):286-92.
Kelly BJ et al. (2016) Composition and dynamics of the respiratory tract microbiome in intubated patients. Microbiome, 4:7.
Marston HD et al. (2016) Antimicrobial Resistance. JAMA, 316(11): 1193-1204.
Martínez-Martínez L, Calvo J (2010) Desarrollo de las resistencias a los antibióticos: causas, consecuencias y su importancia para la salud pública. Enferm Infecc Microbiol Clin, 28(Suppl 4):4-9.
Menis A et al. (2011) Condition of Cleanliness of Surfaces Close to Patients in an Intensive Care Unit. Rev Latino-Am Enfer magem, 19(3):557-64.
Miyaki M et al. (2005) Sequential microbiological monitoring of tracheal aspirates in intubated patients admitted to a pediatric intensive care unit. J Pediatr (Rio J), 81(1):3-4.
Mourani PM et al. (2021) Temporal airway microbiome changes related to ventilator-associated pneumonia in children. Eur Respir J, 57(3):2001829.
Nair GB, Niederman MS (2015) VAP: present understanding and ongoing debates. Intensive Care Med, 41:34-48.
Nante N et al. (2017) Effectiveness of ATP bioluminescence to assess hospital cleaning: a review. J Prev Med Hyg, 58(2):E177-E183.
Nathan C, Cars O (2014) Antibiotic Resistance Problems, Progress, and Prospects. N Engl J Med, 371(19):1761-3.
Pouline M et al. (2017) Airway microbiome research: a modern perspective on surveillance cultures? Ann Transl Med, 5(22):44.
Rosas-Salazar C et al. (2016) Differences in the Nasopharyngeal Microbiome During Acute Respiratory Tract Infection With Human Rhinovirus and Respiratory Syncytial Virus in Infancy. J Infect Dis, 214(12):1924-1928.
Tanner ACR et al. (2018) The Caries Microbiome: Implications for Reversing Dysbiosis. Advances in Dental Research, 29(1):78–85.
The Human Microbiome Project Consortium (2012) Structure, function and diversity of the healthy human microbiome. Nature, 486:207-14.
Wexell C et al. (2016). Antimicrobial Effect of a Single Dose of Amoxicillin on the Oral Microbiota. Clin Implant Dent Relat Res, 18(4):699-706.
White F et al. (2007) A microbiological evaluation of hospital cleaning methods. J Environ Health, 17:285–295.
Wu BG, Segal LN (2018) The Lung Microbiome and Its Role in Pneumonia. Clin Chest Med, 39(4):677-689.
Zakharkina T et al. (2017) The dynamics of the pulmonary microbiome during mechanical ventilation in the intensive care unit and the association with occurrence of pneumonia. Thorax, 72:803–810.