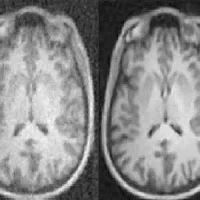
A new deep-learning algorithm can be used to identify and segment tumours in medical images. Developed by AI researchers in Canada, the software makes it possible to automatically analyse several medical imaging modalities, according to a study published in the journal Medical Image Analysis.
“The algorithm makes it possible to automate pre-processing detection and segmentation (delineation) tasks of images, which are currently not done because they are too time-consuming for human beings," explained senior author Samuel Kadoury, a researcher at the University of Montreal Hospital Research Centre (CRCHUM) and professor at Polytechnique Montréal.
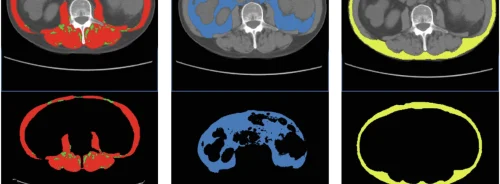
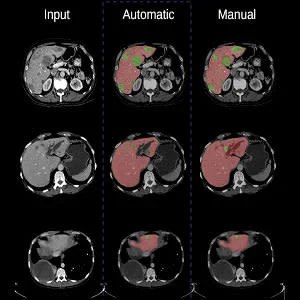
Through a supervised learning process from labelled data inspired by the functioning of the neurons of the brain, the software automatically identifies liver tumours, delineates the contours of the prostate for radiation therapy or makes it possible to count the number of cells at the microscopic level with a performance similar to that of an expert human eye.
The software, Prof. Kadoury said, could be added to visualisation tools to help doctors perform advanced analyses of different medical imaging modalities. "Our model is very versatile – it works for CT liver scan images, magnetic resonance images (MRI) of the prostate and electronic microscopic images of cells,” he added.
Take the example of a patient with liver cancer. Currently, when this patient has a CT scan, the image has to be standardised and normalised before being read by the radiologist. This pre-processing step involves a bit of magic. “You have to adjust the grey shades because the image is often too dark or too pale to distinguish the tumours,” said Dr. An Tang, a radiologist and researcher at the CRCHUM, professor at Université de Montréal and the study’s co-author. “This adjustment with CAD-type computer-aided diagnosis-assistance techniques is not perfect and lesions can sometimes be missed or incorrectly detected. This is what gave us the idea of improving machine vision. The new deep-learning technique eliminates this pre-processing step by modelling the variability observed on a training database.”
The model was developed by the researchers by combining two types of convolutional neural networks that "complemented each another very nicely" to create an image segmentation. The first network takes as an input a raw biomedical data and learns the optimal data normalisation. The second takes the output of the first model and produces segmentation maps,” summarised Michal Drozdzal, the study’s first author, formerly a postdoctoral fellow at Polytechnique and presently research scientist at Facebook AI Research in Montréal.
A neural network is a complex series of computer operations that allows the computer to learn by itself by feeding it a massive number of examples. Convolutional neural networks (CNNs) work a little like our visual cortex by stacking several layers of processing to produce an output result – an image. They can be represented as a pile of building blocks.
The researchers combined two neural networks: a fully convolutional network (FCN) and a fully convolutional residual network (FC-ResNet). They compared the results obtained by their algorithm with other algorithms. “From a visual analysis, we can see that our algorithm performs as well as, if not better than, other algorithms, and is very close to what a human would do if he/she had hours to segment a large number of images. Our algorithm could potentially be used to standardise images from different hospital centres,” asserted Prof. Kadoury.
In cancer imaging, doctors would like to measure the tumour burden, which is the volume of all the tumours in a patient’s body.
“The new algorithm could be used as a tool for detecting and monitoring the tumour burden, to get a much fuller picture of the extent of the disease,” stated Dr. Tang. However, the doctor noted that a number of challenges must be dealt with before these algorithms can be implemented on a large scale. "We’re still in the research and development category. We’ll have to validate it on a large population, in different image-acquisition scenarios, to confirm the robustness of this algorithm,” Dr. Tang added.
Source: Polytechnique Montréal
Image Credit: Eugene Vorontsov
References:
Kadoury S, Drozdza M, Chartrand G et al. (2018) Learning normalized inputs for iterative estimation in medical image segmentation. Medical Image Analysis, February 2018 44: 1-13 https://doi.org/10.1016/j.media.2017.11.005
Latest Articles
Tumours, AI, imaging modalities, Deep-learning technique
A new deep-learning algorithm can be used to identify and segment tumours in medical images. Developed by AI researchers in Canada, the software makes it possible to automatically analyse several medical imaging modalities, according to a study published