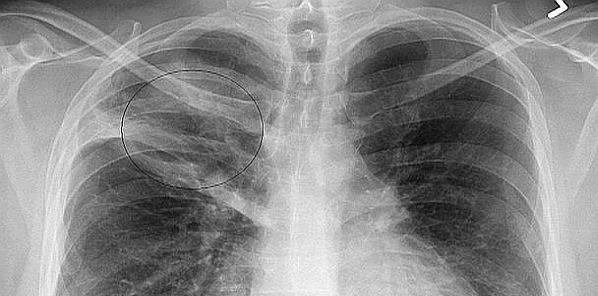
Researchers at George Washington University (GW) in Washington, D.C., have developed a new method for more accurate and rapid identification of bacterial pathogens in patients with pulmonary infections. Next-generation sequencing (NGS) of samples from the sputum of intubated patients, as described in their recently published paper in the Journal of Clinical Microbiology, may enable more focused treatment of pneumonia in the critically ill. This innovation has the potential to reduce healthcare spending and improve patient survival, said the interdisciplinary research team from GW.
It is not uncommon for intensive care unit (ICU) patients to develop ventilator-associated pneumonia and this condition may complicate treatment. Pulmonary infections in critically ill patients often result in significant morbidity, mortality, and additional healthcare costs, according to intensivists.
Currently, ICU patients who experience pneumonia are subjected to broad-spectrum antibiotics. "(This) adds costs, potentially increases the risk of development of antimicrobial resistance, and creates a greater likelihood of an adverse effect attributable to the antibiotics,” said co-author Gary Simon, MD, Ph., Walter G. Ross Professor of Medicine and director of the Division of Infectious Diseases at the GW School of Medicine and Health Sciences (SMHS). “In our paper, we show these methods could improve if we establish a more precise microbiologic cause.”
NGS is the process of determining the DNA sequence of a patient’s genome and microbiome. This genome sequencing technique provides the means to establish a more precise microbiologic cause, according to co-author Timothy McCaffrey, PhD, professor of medicine and director of the Division of Genomic Medicine at GW SMHS. “Through analysing the data provided by the NGS, we were able to identify bacteria not previously identified through standard microbiological methods,” Prof. McCaffrey said.
Interdisciplinary Research Effort
Ian Toma, MD, PhD, MSHS, visiting assistant professor in the Division of Genomic Medicine and Department of Physical Therapy and Health Care Sciences at GW SMHS, developed the NGS procedure using the most advanced sequencing methods available. “It was a challenging proof-of-concept study and a truly interdisciplinary translational research effort that will likely be implemented into clinical practice within the near future,” Dr. Toma said.
NGS data analysis was performed with the help of a bioinformatics tool called “PathoScope", a promising application for identification of pathogens. The tool was developed by a group of researchers led by Keith Crandall, PhD, director of the Computational Biology Institute, a new interdisciplinary research strategic initiative at GW.
Personalised Medicine
“Our tool provides a powerful statistical approach for sifting through NGS data and quickly identifying and characterising pathogens from a patient’s sample,” said Crandall. “This is truly ‘personalised medicine’ as we identify specific strains of bacteria infecting individual patients and provide physicians with targeted information for antibiotic treatments for each individual.”
As technical advances reduce processing and sequencing times, NGS-based methods may ultimately be able to provide physicians with fast, accurate, culture-independent identification of viral, fungal, and bacterial pathogens and their antimicrobial sensitivity profiles.
The interdisciplinary research project was funded in part through a Children’s National Health System and GW Clinical and Translational Science Award grant and by private funding provided by the Abramson Family Foundation, Inc.
Source: GW School of Medicine and Health Sciences
Image Credit: Wikipedia
Currently, ICU patients who experience pneumonia are subjected to broad-spectrum antibiotics. "(This) adds costs, potentially increases the risk of development of antimicrobial resistance, and creates a greater likelihood of an adverse effect attributable to the antibiotics,” said co-author Gary Simon, MD, Ph., Walter G. Ross Professor of Medicine and director of the Division of Infectious Diseases at the GW School of Medicine and Health Sciences (SMHS). “In our paper, we show these methods could improve if we establish a more precise microbiologic cause.”
NGS is the process of determining the DNA sequence of a patient’s genome and microbiome. This genome sequencing technique provides the means to establish a more precise microbiologic cause, according to co-author Timothy McCaffrey, PhD, professor of medicine and director of the Division of Genomic Medicine at GW SMHS. “Through analysing the data provided by the NGS, we were able to identify bacteria not previously identified through standard microbiological methods,” Prof. McCaffrey said.
Interdisciplinary Research Effort
Ian Toma, MD, PhD, MSHS, visiting assistant professor in the Division of Genomic Medicine and Department of Physical Therapy and Health Care Sciences at GW SMHS, developed the NGS procedure using the most advanced sequencing methods available. “It was a challenging proof-of-concept study and a truly interdisciplinary translational research effort that will likely be implemented into clinical practice within the near future,” Dr. Toma said.
NGS data analysis was performed with the help of a bioinformatics tool called “PathoScope", a promising application for identification of pathogens. The tool was developed by a group of researchers led by Keith Crandall, PhD, director of the Computational Biology Institute, a new interdisciplinary research strategic initiative at GW.
Personalised Medicine
“Our tool provides a powerful statistical approach for sifting through NGS data and quickly identifying and characterising pathogens from a patient’s sample,” said Crandall. “This is truly ‘personalised medicine’ as we identify specific strains of bacteria infecting individual patients and provide physicians with targeted information for antibiotic treatments for each individual.”
As technical advances reduce processing and sequencing times, NGS-based methods may ultimately be able to provide physicians with fast, accurate, culture-independent identification of viral, fungal, and bacterial pathogens and their antimicrobial sensitivity profiles.
The interdisciplinary research project was funded in part through a Children’s National Health System and GW Clinical and Translational Science Award grant and by private funding provided by the Abramson Family Foundation, Inc.
Source: GW School of Medicine and Health Sciences
Image Credit: Wikipedia
Latest Articles
ICU, ventilator associated pneumonia, next-generation sequencing, NGS, critically ill
Researchers at George Washington University (GW) in Washington, D.C., have developed a new method for more accurate and rapid identification of bacterial p...