HealthManagement, Volume 20 - Issue 10, 2020
Machine learning and radiomics may lead to personalised management and treatment of glioma patients, hopefully improving quality of life and survival.
Key Points
- In the past few years, multiple radiomics-based tools have been introduced in the field of neuro-oncology and in particular for imaging of brain gliomas.
- Radiogenomics may evaluate molecular expression of brain gliomas in the pre-surgical setting.
- Radiomics may predict the grading of brain gliomas without the need of invasive procedures.
- Prediction of prognosis of brain gliomas may be helped by radiomics in the near future.
Recently, there has been a growing interest in medicine for radiomics as it could have multiple applications in numerous fields (Cuocolo et al. 2019). Machine Learning (ML) is a subfield of radiomics which could be used to solve problems through the use of algorithms that dynamically learn from available data, without the need of prior explicit programming. It can be classified in three types based on the type of learning: supervised (if there is an available ground truth guiding the training process), unsupervised and reinforcement learning (when the algorithm learns from reinforcement given by a dynamic environment) (Choy et al. 2018). Deep Learning (DL) is a complex ML model consisting of multi-layered networks, each composed by a web of nodes. Therefore, it allows high-level abstraction of data and good performance (Cuocolo and Ugga 2018). Radiogenomics, also known as imaging genomics, is a field of radiomics which identifies relationships between tumour genomic characteristics and imaging phenotypes (Zhou et al. 2018). By far, radiology is the field of medicine with the most FDA-approved radiomics-based tools, in particular in the subfields of neuro-oncology (Cuocolo et al. 2020).
Gliomas represent the most common primary brain tumours, with an incidence ranging from 0.2-4.8 per 100,000 (depending on the specific subtype) and being more frequent in adults between 45 and 65 years (Kickingereder and Bisdas 2019). There are four grades according to the World Health Organization (WHO) histopathological classification: grades I and II are low grade gliomas whereas grade III and IV are considered high grade. After the 2016 WHO classification of central nervous system tumours, they have also been classified based on molecular parameters, as they strongly influence survival. In particular, the mutation status of Isocitrate dehydrogenase (IDH), the expression of ATRX, 1p/19q codeletion, K27 M mutations in thegene H3F3A and MGMT methylation status have been strongly associated with survival outcome (Louis et al. 2016).
Radiomics has proved to be useful in the imaging of brain gliomas, in particular in pre-operative and intra-operative planning, histopathologic diagnosis, post-treatment follow-up and outcome prediction (Sotoudeh et al. 2019). Hopefully, in the near future there will be ML-tools able to accurately evaluate tumoural gene expression, grading and survival prediction in the pre-surgical setting.
Radiogenomics and Tumoural Gene Expression
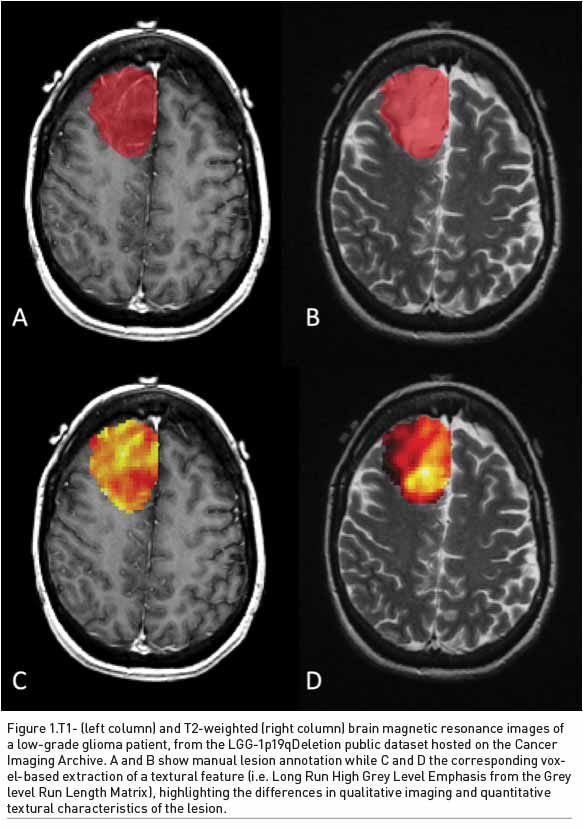
Prediction of molecular status of brain gliomas in the pre-surgical setting has become crucial to increase the efficacy of treatment. In this setting, radiogenomics may play a crucial role (Figure 1).
Buda et al. (2020) developed a DL tool able to predict genomic subtypes of lower-grade gliomas (WHO grade II and III) using magnetic resonance (MR) images. They collected 110 patients from five institutions and the DL model achieved area under the receiver operating characteristic curve (AUC) of 0.730 for the prediction of the tumour genomics (Buda et al. 2020).
In particular, the evaluation of IDH mutation status is extremely important for the characterisation of gliomas and treatment choice. V-Net is a tool based on convolutional neural network, a subtype of DL, that uses 3D-layers for the automatic segmentation of medical images. It has proved to be efficient in the evaluation and segmentation of Glioblastoma Multiforme (GBM) and peritumoural oedema on T2-weighted MR images and in the prediction of the mutation status of IDH1 (AUC=0.86) (Choi et al. 2020a).
Qian et al. (2018) created a radiomics signature based on T2-weighted MR images of low-grade gliomas which integrated radiomics and radiogenomics. It showed good correlation with hypoxia, angiogenesis, apoptosis and cell proliferation and could predict the presence of unfavourable gene expression (Qian et al. 2018). Furthermore, it showed good correlation with prognosis.
Finally, MR could help in the evaluation of MGMT methylation, an important biomarker for the prediction of response to alkylating chemotherapy. Li et al. created a radiomics model which could predict MGMT methylation status in GBM from multiregional and multiparametric MR images. The primary cohort was composed by 133 patients whereas the validation cohort was composed by 60 patients. The radiomics model reached an AUC of 0.88 ( Li et al. 2018).
Prediction of Grading in the Pre-surgical Setting
Non-invasive tools for prediction of grading of brain gliomas could be extremely useful. Therefore, multiple radiomics-based tools have been developed, though they still require adequate validation for daily use in clinical practice.
For instance, Takahashi et al. created a ML algorithm based on MR images which can predict the grading of glioma with high accuracy. In particular, features extracted from apparent diffusion coefficient reached an AUC of 0.95.
On the other hand, Li et al. (2020) developed a radiomics tool based on T2 and FLAIR images for prediction of specific immunohistochemical biomarkers of gliomas. It was able to estimate the percentage of presence of Ki-67, d-100, vimentin and CD34, which are associated with tumour proliferation, malignancy, therapeutic effectiveness, invasion and angiogenesis. The AUC ranged from 0.713 to 0.923, depending on this specific biomarker (Li et al. 2020).
DL has proved to be useful in this setting. A DL-based classification trained on a cohort of 121 patients, showed high accuracy (AUC=0.987) in the estimation of glioma grading from brain MR (Çinarer, et al. 2020). Similarly, Chen et al. created a tool bases on convolutional neural network, a subtype of DL, for automatic segmentation and grading evaluation of gliomas. It used a training cohort of 220 high-grade gliomas an 54 low-grade gliomas, reaching an accuracy of 91.27% (Chen et al. 2018).
Prognosis and Survival Prediction
Radiomics has been used to predict overall survival of patients with gliomas. Baid et al. (2020) created a radiomics tool based on imaging features extracted from FLAIR and contrast enhanced T1 of gliomas to predict overall survival. It achieved high accuracy in the training, validation and test datasets (0.695, 0.571 and 0.558 respectively) (Baid et al. 2020). Similarly, Shboul et al. created a DL tool based on MR for automated segmentation and classification of GBM and survival prediction with an accuracy of 0.73 for training datasets and 0.68 for validation cohort (Shboul et al. 2019).
Prediction of survival of ML tools increases when imaging features are combined with conventional clinical prognostic factors, such as sex, age, Karnofsky performance score (KPS), tumour location, tumour volume, MGMT promoter status and extent of surgery. For instance, radiomics based on multiparametric-MR showed high accuracy in prediction of overall survival and performance status in GBM when combined with conventional clinical and genetic prognostic models (Choi et al. 2020b). Park et al. developed a radiomics model combining multiparametric MR and clinical predictors for prediction of prognosis in newly diagnosed GBM. In particular, data from conventional, diffusion- and perfusion-weighted-MR taken from 216 patients were combined withclinical and genetic predictors and it outperformed models based on clinical predictors alone with a C-index of 0.74 (Park et al. 2020).
An important issue is to create radiomics tool that can be used in different institution with high accuracy and reproducibility. In particular, Suter et al. developed a ML tool for prediction of overall survival in GBM that showed robustness and accuracy when used to unseen multi-centre data (Suter et al. 2020).
Conclusion
ML showed promising results in the imaging of gliomas, thus there will probably be an increasing number of radiomics-based tools in this field, in particular for pre-surgical evaluation of tumoural gene expression, grading and prediction of prognosis. This will lead to a more personalised management and treatment of patients with gliomas, hopefully improving quality of life and survival.
Conflict of Interest
None.
References:
Baid U et al. (2020) Overall Survival Prediction in Glioblastoma With Radiomic Features Using Machine Learning. Frontiers in Computational Neuroscience, 1–9.
Buda M et al. (2020) Deep Radiogenomics of Lower-Grade Gliomas: Convolutional Neural Networks Predict Tumor Genomic Subtypes Using MR Images. Radiology: Artificial Intelligence, 2(1): e180050.
Chen W et al. (2018) Computer-Aided Grading of Gliomas Combining Automatic Segmentation and Radiomics. International Journal of Biomedical Imaging, 1–11.
Choi Y et al. (2020a) IDH1 Mutation Prediction Using MR-Based Radiomics in Glioblastoma: Comparison between Manual and Fully Automated Deep Learning-Based Approach of Tumor Segmentation. European Journal of Radiology, 128:109031.
Choi Y et al. (2020b) Radiomics May Increase the Prognostic Value for Survival in Glioblastoma Patients When Combined with Conventional Clinical and Genetic Prognostic Models. European Radiology.
Choy G et al. (2018) Current Applications and Future Impact of Machine Learning in Radiology. Radiology 288(2): 318–28.
Çinarer G et al. (2020) Prediction of Glioma Grades Using Deep Learning with Wavelet Radiomic Features. Applied Sciences 10(18): 6296.
Cuocolo R et al. (2019) Current Applications of Big Data and Machine Learning in Cardiology. Journal of geriatric cardiology, 16(8): 601–7.
Cuocolo R et al. (2020) Machine Learning in Oncology: A Clinical Appraisal. Cancer Letters 481: 55–62.
Cuocolo R, Ugga L (2018) Imaging Applications of Artificial Intelligence. HealthManagement.org The Journal 18(6).
Kickingereder P, Sotirios B (2019) Glial Tumors and Primary CNS Lymphoma. In Clinical Neuroradiology, Cham: Springer International Publishing, 1051–74.
Li J et al. (2020) High-Order Radiomics Features Based on T2 FLAIR MRI Predict Multiple Glioma Immunohistochemical Features: A More Precise and Personalized Gliomas Management” ed. Alessandro Weisz. PLOS ONE 15(1): e0227703.
Li, Z-C et al. (2018) Multiregional Radiomics Features from Multiparametric MRI for Prediction of MGMT Methylation Status in Glioblastoma Multiforme: A Multicentre Study.” European Radiology 28(9): 3640–50.
Louis DN. et al.( 2016) The 2016 World Health Organization Classification of Tumors of the Central Nervous System: A Summary. Acta Neuropathologica 131(6): 803–20.
Park JE et al. (2020)“Radiomics Prognostication Model in Glioblastoma Using Diffusion- and Perfusion-Weighted MRI. Scientific Reports 10(1): 4250.
Qian Z et al. (2018) Radiogenomics of Lower-Grade Gliomas: A Radiomic Signature as a Biological Surrogate for Survival Prediction. Aging 10(10): 2884–99.
Shboul ZA et al. (2019) Feature-Guided Deep Radiomics for Glioblastoma Patient Survival Prediction. Frontiers in Neuroscience 13:1–17.
Sotoudeh H et al. (2019) Artificial Intelligence in the Management of Glioma: Era of Personalized Medicine. Frontiers in Oncology, 9:1–11.
Suter Y et al. (2020) Radiomics for Glioblastoma Survival Analysis in Pre-Operative MRI: Exploring Feature Robustness, Class Boundaries, and Machine Learning Techniques. Cancer Imaging 20(1): 55.
Zhou M et al. (2018) Radiomics in Brain Tumor: Image Assessment, Quantitative Feature Descriptors, and Machine-Learning Approaches. American Journal of Neuroradiology 39(2): 208–16.





