Research scientists at
the University of Bristol have used computer simulations to show how bacteria
destroy antibiotics. This could be a major breakthrough and could help in the development of drugs that can effectively tackle bacterial infections in the future. The research
team focused on the role of enzymes in the bacteria and how they split the
structure of the antibiotic, making it bacteria-resistant and essentially
ineffective.
The research was carried out by academics and postgraduate and undergraduate students at the School of Cellular and Molecular Medicine and the School of Chemistry. It was funded by the Engineering and Physical Sciences Research Council (EPSRC). The new findings have been published in Chemical Communications and can offer insights for scientists, enabling them to develop new antibiotics that have a lower risk of resistance. They can also help identify and use the best medicines for specific outbreaks.
The research team used prize-winning technique called quantum mechanics/molecular mechanics simulations (QM/MM). The technique was developed by 2013 Nobel Prize winner Martin Karplus, Michael Levitt and Arieh Warshel.
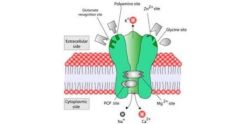
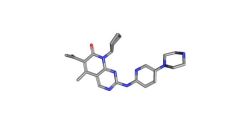
The QM/MM technique allowed the researchers to gain a molecular-level insight into how the beta-lactamase enzyme reacts to antibiotics. The primary area of interest of the research team is to study the growing resistance to carbapenems, generally considered to be the last-resort antibiotics for several bacterial infections and super bugs such as E. coli. This growing resistance is making many bacterial infections difficult to treat or even untreatable and has made even minor infections dangerous and possibly fatal.
The computer simulations showed that the most important step occurs when the enzyme spits out the broken-down antibiotic. A bacterium becomes resistant when the spitting out process happens too quickly, since it then goes on chewing up antibiotics. If the process is slow, the enzyme gets clogged and is unable to break the antibiotic anymore. The bacterium dies and does not become resistant. The spitting out speed depends on the height of the energy barrier for the reaction. If the barrier is low, the rate of spitting-out is quicker and if the barrier is high, the rate of spitting-out is slow.
According to Professor Adrian Mulholland, from Bristol University’s School of Chemistry, “We've shown that we can use computer simulations to identify which enzymes break down and spit out carbapenems quickly and those that do it only slowly. This means that these simulations can be used in future to test enzymes and predict and understand resistance. We hope that this will identify how they act against different drugs – a useful tool in developing new antibiotics and helping to choose which drugs might be best for treating a particular outbreak.”
Source: Bristol.ac.uk
Image credit: Flickr.com