An multinational team of researchers led by Tomas Ekeberg from Uppsala University in Sweden has demonstrated that it is possible to determine the 3D structure of a virus by combining several diffraction patterns obtained from a sequence of randomly oriented single viruses. The technique does not depend on crystallisation, which is how scientists in the past have examined molecular structures. Instead, the team utilised Stanford’s Linac Coherent Light Source (LCLS), one of only two hard x-ray free-electron lasers (XFELs) in operation worldwide.
X-ray crystallography has helped researchers to examine the structure of biological molecules such as nucleic acids and proteins since the 1950s, but that method cannot work for molecules which cannot be crystallised, such as “membrane proteins” which interact with biological membranes. These proteins play a critical role in immune response. The new technique therefore has the potential to image pathogenic viruses including herpes, HIV and influenza.
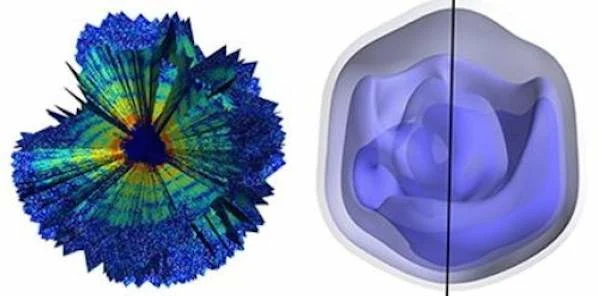
The team of scientists conducted its research using the mimivirus, a giant virus that is thought to cause certain forms of pneumonia. The researchers injected single viruses into the intense beam of the LCLS, recording approximately 200 diffraction patterns from individual viruses. An optimised version of the expand, maximise and compress algorithm allowed the team to validate, based on the diffraction patterns, the relative orientation of the particles within each recording so that they could generate three-dimensional images of the mimivirus.
If the structure of non-crystallisable biomolecules can be rebuilt, the implications for biology would be profound. This research demonstrates the possibility of deriving 3D structures of molecules from the diffraction data of hundreds of individual samples, by assembling the combined data into diffraction patterns.
The relatively large size of the mimivirus made it a suitable choice for use in the study. Still, even the mimivirus is only one tenth the width of a human hair, and the strength of the x-ray beams destroy the molecules that are being imaged. Eventually, the reseachers hope to perfect their method so that images of living bacteria and viruses can be rendered. The current 125 nanometre spatial resolution will need to be improved upon in order to tackle smaller molecules such as proteins, but these findings are a promising step in the right direction.
Source: American Physical Society
Image Credit: Tomas Ekeberg et al., Physical Review Letters